|
To understand how the Measure Your Gradient methodology works, you first have to understand how gradient retention times are calculated from isocratic retention measurements.
Suppose you measured the isocratic retention of a compound over a range of mobile phase compositions...
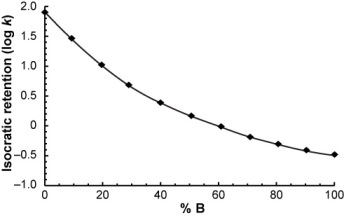
...and you want to calculate its retention time in the following gradient.
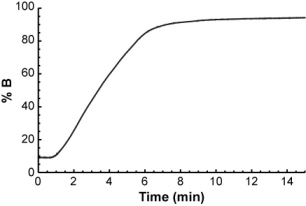
The easiest way to do it is to think of the gradient as a series of very short isocratic steps, each with a certain mobile phase composition.
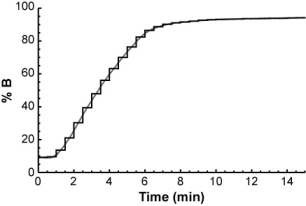
For each isocratic step, you could determine the compound's retention factor (from the isocratic retention vs. % B relationship above) and from that, calculate how far it will travel through the column during the step. After a number of isocratic steps, the compound will eventually have traveled the entire length of the column.
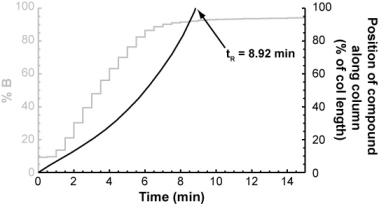
The number of isocratic steps it took to elute the compound multiplied by the length of each step gives its gradient retention time.
The Measure Your Gradient methodology uses the same basic approach, but in reverse. Instead of calculating gradient retention times from a measured gradient profile, it back-calculates what the gradient profile must have been to give a set of measured gradient retention times.
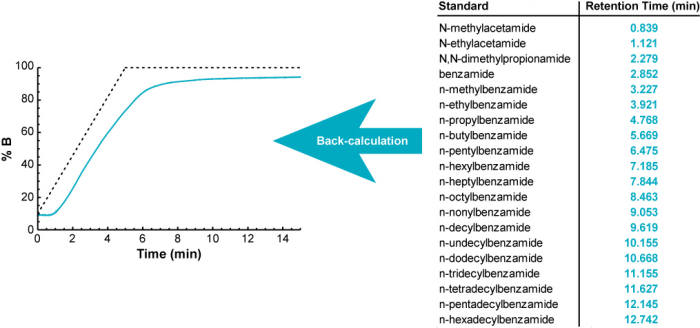
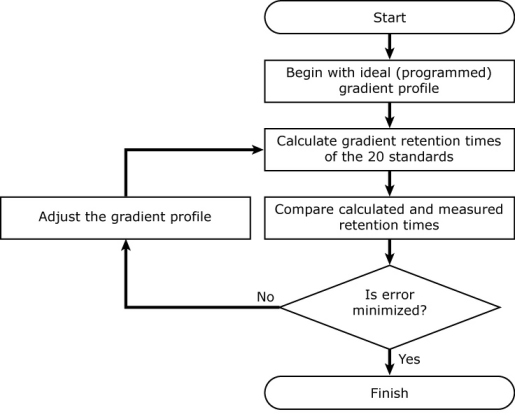
It uses an iterative process to back-calculate the gradient profile. It begins with the ideal (programmed) gradient profile, calculates retention times for each of 20 standards (using their known isocratic retention behavior; see below), and compares them with their measured retention times. Then it makes a small adjustment to the gradient profile, recalculates retention times for all 20 compounds, and checks to see if the accuracy of the calculated retention times improved. If so, it keeps the change, otherwise it discards it. It continues making small adjustments to the gradient profile until the difference between the measured and calculated retention times is minimized.
|
